Abstract
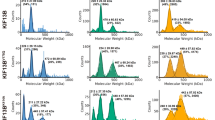
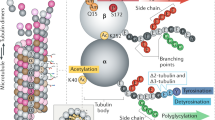
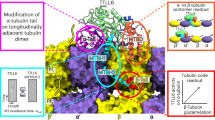
The ‘tubulin-code’ hypothesis proposes that different tubulin genes or post-translational modifications (PTMs), which mainly confer variation in the carboxy-terminal tail (CTT), result in unique interactions with microtubule-associated proteins for specific cellular functions. However, the inability to isolate distinct and homogeneous tubulin species has hindered biochemical testing of this hypothesis. Here, we have engineered 25 α/β-tubulin heterodimers with distinct CTTs and PTMs and tested their interactions with four different molecular motors using single-molecule assays. Our results show that tubulin isotypes and PTMs can govern motor velocity, processivity and microtubule depolymerization rates, with substantial changes conferred by even single amino acid variation. Revealing the importance and specificity of PTMs, we show that kinesin-1 motility on neuronal β-tubulin (TUBB3) is increased by polyglutamylation and that robust kinesin-2 motility requires detyrosination of α-tubulin. Our results also show that different molecular motors recognize distinctive tubulin ‘signatures’, which supports the premise of the tubulin-code hypothesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Nogales, E. Structural insight into microtubule function. Annu. Rev. Biophys. Biomol. Struct. 30, 397–420 (2001).
Amos, L. A. & Schlieper, D. Microtubules and maps. Adv. Protein Chem. 71, 257–298 (2005).
Ludueña, R. F. & Banerjee, A. in Role Microtubules Cell Biol. Neurobiol. Oncol. (ed Md, T. F.) 123–175 (Humana Press, 2008).
Banerjee, A. et al. A monoclonal antibody against the type II isotype of β-tubulin. Preparation of isotypically altered tubulin. J. Biol. Chem. 263, 3029–3034 (1988).
Burgoyne, R. D., Cambray-Deakin, M. A., Lewis, S. A., Sarkar, S. & Cowan, N. J. Differential distribution of β-tubulin isotypes in cerebellum. EMBO J. 7, 2311–2319 (1988).
Raff, E. C., Fackenthal, J. D., Hutchens, J. A., Hoyle, H. D. & Turner, F. R. Microtubule architecture specified by a β-tubulin isoform. Science 275, 70–73 (1997).
Leandro-García, L. J. et al. Hematologic β-tubulin VI isoform exhibits genetic variability that influences paclitaxel toxicity. Cancer Res. 72, 4744–4752 (2012).
Wang, D., Villasante, A., Lewis, S. A. & Cowan, N. J. The mammalian β-tubulin repertoire: hematopoietic expression of a novel, heterologous β-tubulin isotype. J. Cell Biol. 103, 1903–1910 (1986).
Cederquist, G. Y. et al. An inherited TUBB2B mutation alters a kinesin-binding site and causes polymicrogyria, CFEOM and axon dysinnervation. Hum. Mol. Genet. 21, 5484–5499 (2012).
Tischfield, M. A. et al. Human TUBB3 mutations perturb microtubule dynamics, kinesin interactions, and axon guidance. Cell 140, 74–87 (2010).
Tischfield, M. A., Cederquist, G. Y., Gupta, M. L. Jr & Engle, E. C. Phenotypic spectrum of the tubulin-related disorders and functional implications of disease-causing mutations. Curr. Opin. Genet. Dev. 21, 286–294 (2011).
Tischfield, M. A. & Engle, E. C. Distinct α- and β-tubulin isotypes are required for the positioning, differentiation and survival of neurons: new support for the ‘multi-tubulin’ hypothesis. Biosci. Rep. 30, 319–330 (2010).
Breuss, M. et al. Mutations in the β-tubulin gene TUBB5 cause microcephaly with structural brain abnormalities. Cell Rep. 2, 1554–1562 (2012).
L’Hernault, S. W. & Rosenbaum, J. L. Chlamydomonas α-tubulin is posttranslationally modified by acetylation on the ε-amino group of a lysine. Biochemistry 24, 473–478 (1985).
Janke, C. & Bulinski, J. C. Post-translational regulation of the microtubule cytoskeleton: mechanisms and functions. Nat. Rev. Mol. Cell Biol. 12, 773–786 (2011).
Garnham, C. P. & Roll-Mecak, A. The chemical complexity of cellular microtubules: tubulin post-translational modification enzymes and their roles in tuning microtubule functions. Cytoskelet. Hoboken NJ 69, 442–463 (2012).
Peris, L. et al. Motor-dependent microtubule disassembly driven by tubulin tyrosination. J. Cell Biol. 185, 1159–1166 (2009).
Konishi, Y. & Setou, M. Tubulin tyrosination navigates the kinesin-1 motor domain to axons. Nat. Neurosci. 12, 559–567 (2009).
Dunn, S. et al. Differential trafficking of Kif5c on tyrosinated and detyrosinated microtubules in live cells. J. Cell Sci. 121, 1085–1095 (2008).
Liao, G. & Gundersen, G. G. Kinesin is a candidate for cross-bridging microtubules and intermediate filaments. Selective binding of kinesin to detyrosinated tubulin and vimentin. J. Biol. Chem. 273, 9797–9803 (1998).
Akhmanova, A. & Steinmetz, M. O. Tracking the ends: a dynamic protein network controls the fate of microtubule tips. Nat. Rev. Mol. Cell Biol. 9, 309–322 (2008).
Eddé, B. et al. Posttranslational glutamylation of α-tubulin. Science 247, 83–85 (1990).
Audebert, S. et al. Reversible polyglutamylation of α- and β-tubulin and microtubule dynamics in mouse brain neurons. Mol. Biol. Cell 4, 615–626 (1993).
Janke, C. et al. Tubulin polyglutamylase enzymes are members of the TTL domain protein family. Science 308, 1758–1762 (2005).
Wloga, D. & Gaertig, J. Post-translational modifications of microtubules. J. Cell Sci. 123, 3447–3455 (2010).
Van Dijk, J. et al. A targeted multienzyme mechanism for selective microtubule polyglutamylation. Mol. Cell 26, 437–448 (2007).
Audebert, S. et al. Developmental regulation of polyglutamylated α- and β-tubulin in mouse brain neurons. J. Cell Sci. 107 (Pt 8), 2313–2322 (1994).
Redeker, V. Mass spectrometry analysis of C-terminal posttranslational modifications of tubulins. Methods Cell Biol. 95, 77–103 (2010).
Gaertig, J. & Wloga, D. Ciliary tubulin and its post-translational modifications. Curr. Top. Dev. Biol. 85, 83–113 (2008).
Lacroix, B. et al. Tubulin polyglutamylation stimulates spastin-mediated microtubule severing. J. Cell Biol. 189, 945–954 (2010).
Bonnet, C. et al. Differential binding regulation of microtubule-associated proteins MAP1A, MAP1B, and MAP2 by tubulin polyglutamylation. J. Biol. Chem. 276, 12839–12848 (2001).
Fulton, C. & Simpson, P. A. Selective synthesis and utilization of flagellar tubulin. The multitubulin hypothesis. Cell Motil. 3, 987–1005 (1976).
Verhey, K. J. & Gaertig, J. The tubulin code. Cell Cycle Georget. Tex 6, 2152–2160 (2007).
Serrano, L., Avila, J. & Maccioni, R. B. Controlled proteolysis of tubulin by subtilisin: localization of the site for MAP2 interaction. Biochemistry 23, 4675–4681 (1984).
Redeker, V., Melki, R., Promé, D., Le Caer, J. P. & Rossier, J. Structure of tubulin C-terminal domain obtained by subtilisin treatment. The major α and β tubulin isotypes from pig brain are glutamylated. FEBS Lett. 313, 185–192 (1992).
Lobert, S., Hennington, B. S. & Correia, J. J. Multiple sites for subtilisin cleavage of tubulin: effects of divalent cations. Cell Motil. Cytoskeleton 25, 282–297 (1993).
Johnson, V., Ayaz, P., Huddleston, P. & Rice, L. M. Design, overexpression, and purification of polymerization-blocked yeast αβ-tubulin mutants. Biochemistry 50, 8636–8644 (2011).
Sindelar, C. V. & Downing, K. H. An atomic-level mechanism for activation of the kinesin molecular motors. Proc. Natl Acad. Sci. USA 107, 4111–4116 (2010).
Redwine, W. B. et al. Structural basis for microtubule binding and release by dynein. Science 337, 1532–1536 (2012).
Gigant, B. et al. Structure of a kinesin-tubulin complex and implications for kinesin motility. Nat. Struct. Mol. Biol. 20, 1001–1007 (2013).
Thorn, K. S., Ubersax, J. A. & Vale, R. D. Engineering the processive run length of the kinesin motor. J. Cell Biol. 151, 1093–1100 (2000).
Hirokawa, N., Noda, Y., Tanaka, Y. & Niwa, S. Kinesin superfamily motor proteins and intracellular transport. Nat. Rev. Mol. Cell Biol. 10, 682–696 (2009).
Kardon, J. R. & Vale, R. D. Regulators of the cytoplasmic dynein motor. Nat. Rev. Mol. Cell Biol. 10, 854–865 (2009).
Reck-Peterson, S. L. et al. Single-molecule analysis of dynein processivity and stepping behavior. Cell 126, 335–348 (2006).
Trokter, M., Mücke, N. & Surrey, T. Reconstitution of the human cytoplasmic dynein complex. Proc. Natl Acad. Sci. USA 109, 20895–20900 (2012).
Wang, Z. & Sheetz, M. P. The C-terminus of tubulin increases cytoplasmic dynein and kinesin processivity. Biophys. J. 78, 1955–1964 (2000).
Cooper, J. R., Wagenbach, M., Asbury, C. L. & Wordeman, L. Catalysis of the microtubule on-rate is the major parameter regulating the depolymerase activity of MCAK. Nat. Struct. Mol. Biol. 17, 77–82 (2010).
Hertzer, K. M. & Walczak, C. E. The C-termini of tubulin and the specific geometry of tubulin substrates influence the depolymerization activity of MCAK. Cell Cycle Georget. Tex 7, 2727–2737 (2008).
Rüdiger, M., Plessman, U., Klöppel, K. D., Wehland, J. & Weber, K. Class II tubulin, the major brain β tubulin isotype is polyglutamylated on glutamic acid residue 435. FEBS Lett. 308, 101–105 (1992).
Westermann, S. & Weber, K. Post-translational modifications regulate microtubule function. Nat. Rev. Mol. Cell Biol. 4, 938–947 (2003).
Okada, Y. & Hirokawa, N. Mechanism of the single-headed processivity: diffusional anchoring between the K-loop of kinesin and the C terminus of tubulin. Proc. Natl Acad. Sci. USA 97, 640–645 (2000).
Lakämper, S. & Meyhöfer, E. The E-hook of tubulin interacts with kinesin’s head to increase processivity and speed. Biophys. J. 89, 3223–3234 (2005).
Skiniotis, G. et al. Modulation of kinesin binding by the C-termini of tubulin. EMBO J. 23, 989–999 (2004).
Derr, N. D. et al. Tug-of-war in motor protein ensembles revealed with a programmable DNA origami scaffold. Science 338, 662–665 (2012).
Rai, A. K., Rai, A., Ramaiya, A. J., Jha, R. & Mallik, R. Molecular adaptations allow dynein to generate large collective forces inside cells. Cell 152, 172–182 (2013).
O’Hagan, R. et al. The tubulin deglutamylase CCPP-1 regulates the function and stability of sensory cilia in C. elegans. Curr. Biol. 21, 1685–1694 (2011).
Rogowski, K. et al. A family of protein-deglutamylating enzymes associated with neurodegeneration. Cell 143, 564–578 (2010).
Imanishi, M., Endres, N. F., Gennerich, A. & Vale, R. D. Autoinhibition regulates the motility of the C. elegans intraflagellar transport motor OSM-3. J. Cell Biol. 174, 931–937 (2006).
Hammond, J. W., Blasius, T. L., Soppina, V., Cai, D. & Verhey, K. J. Autoinhibition of the kinesin-2 motor KIF17 via dual intramolecular mechanisms. J. Cell Biol. 189, 1013–1025 (2010).
Schatz, P. J., Georges, G. E., Solomon, F. & Botstein, D. Insertions of up to 17 amino acids into a region of α-tubulin do not disrupt function in vivo. Mol. Cell Biol. 7, 3799–3805 (1987).
Uchimura, S. et al. Identification of a strong binding site for kinesin on the microtubule using mutant analysis of tubulin. EMBO J. 25, 5932–5941 (2006).
Carter, A. P., Cho, C., Jin, L. & Vale, R. D. Crystal structure of the dynein motor domain. Science 331, 1159–1165 (2011).
Tomishige, M. & Vale, R. D. Controlling kinesin by reversible disulfide cross-linking. Identifying the motility-producing conformational change. J. Cell Biol. 151, 1081–1092 (2000).
Yildiz, A., Tomishige, M., Vale, R. D. & Selvin, P. R. Kinesin walks hand-over-hand. Science 303, 676–678 (2004).
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer control of microscopes using μManager. Curr. Protoc. Mol. Biol. Ed. Frederick M Ausubel Al Ch.14, Unit14.20 (2010).
Acknowledgements
The authors thank N. Stuurman for microscopy assistance, M. Tanenbaum for help in lentiviral expression and members of the R.D.V. laboratory for comments on the manuscript. R.D.V. is a Howard Hughes Medical Institute investigator and M.S. is a Human Frontiers Science Program—Long-term fellow (LT-000120/2009). L.M.R. is supported by NSF MCB-1054947. This work received support from an NIH grant (38499).
Author information
Authors and Affiliations
Contributions
M.S. and R.D.V. conceived the project. M.S. performed the experiments, analysed the data and wrote the paper. L.M.R. developed and tested the internal His-tagged tubulin and advised in this study. R.D.V. supervised the work and wrote the paper. All authors discussed and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Conservation of microtubule lattice residues.
Sequence conservation of microtubule lattice surface residues of α- (dark grey) and β- (light grey) tubulin. The surface residues are coloured according their conservation between yeast and human tubulin isotypes (red, 100% conservation).
Supplementary Figure 2
Sequence alignment of human α-tubulin isotypes and yeast TUB1, highlighting the conservation of surface residue that are within 6 Å from the kinesin and dynein microtubule binding interfaces, as defined by crystal structure of kinesin-tubulin and cryo-EM structured of dynein-microtubule complexes. This alignment reveals the extremely high degree of conservation of the core compared with the CTTs (see ??a, ??a), with the few cases of substitutions mostly involving similar amino acids.
Supplementary Figure 3
Sequence alignment of human α-tubulin isotypes and yeast TUB2, as indicated in Supplementary Figure 2.
Supplementary Figure 4 Sequence alignment of human α-tubulin isotypes and yeast TUB2, as indicated in Supplementary Figure 2.
a. The site of the His-6 tag in the luminal loop of α-tubulin (light grey) as illustrated. b. Sequence alignment of α-tubulin (yeast TUB1, amino acids 31-68) from different organisms (HS; Human, SS; Sus scrofa, DM; Drosophila melanogaster, SU, Sea Urchin, CE; Caenorhabditis elegans, SC; Saccharomyces cerevisiae). The His6-tag was introduced in between P43 and K44; an insertion of 17 residues in this non-conserved loop does not disrupt tubulin function in yeast60 (see Online Methods). c. SDS-PAGE (left) of the purified recombinant yeast α-int-His/β-tubulin; the DIC image (right) shows microtubules assembled from this tubulin.
Supplementary Figure 5
Sequence alignment of human and Chinese hamster MCAK (kinesin-13) proteins, the motor domain is highlighted in grey. The sequence identity of full length and motor domain are ∼93% and ∼96% respectively.
Supplementary Figure 6 Specificity of 10E maleimide towards cysteine at CTTs.
SDS-PAGE of cross-linked products from left to right; Untreated TUBA1A-E542C/TUBB2-E435C; TUBA1A-E542C/TUBB2-E435C + 10E maleimide peptide; untreated TUBA1A/TUBB2 cysteine light mutants (CLM); TUBA1A/TUBB2-CLM + 10E maleimide peptide.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1887 kb)
Supplementary Table 1
Supplementary Information (XLSX 69 kb)
Rights and permissions
About this article
Cite this article
Sirajuddin, M., Rice, L. & Vale, R. Regulation of microtubule motors by tubulin isotypes and post-translational modifications. Nat Cell Biol 16, 335–344 (2014). https://doi.org/10.1038/ncb2920
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2920
This article is cited by
-
Novel variants in TUBB8 gene cause multiple phenotypic abnormalities in human oocytes and early embryos
Journal of Ovarian Research (2023)
-
Tubulin engineering by semi-synthesis reveals that polyglutamylation directs detyrosination
Nature Chemistry (2023)
-
Taxol acts differently on different tubulin isotypes
Communications Biology (2023)
-
Control of motor landing and processivity by the CAP-Gly domain in the KIF13B tail
Nature Communications (2023)
-
Regulatory mechanisms of the dynein-2 motility by post-translational modification revealed by MD simulation
Scientific Reports (2023)