Abstract
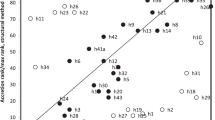
The expansion segments within the eukaryote nuclear 23S-like ribosomal RNA molecule are now well characterized in many diverse organisms. A different base compositional bias, a higher propensity for size variability, and an increased evolutionary rate distinguish these regions from the universally conserved “core” regions of the molecule. In addition, some expansion segments of higher eukaryotes exhibit significant sequence simplicity which is hypothesized to occur by slippage-mediated mutational processes. We describe the discovery of extreme size variation of the D3 expansion segment in the crustacean order Isopoda. Among 11 species D3 varies in size from 180 to 518 nucleotides but maintains a homologous secondary structure. The D3 size is significantly positively correlated to relative simplicity factor (RSF), indicating that growth is most likely by insertion of simple sequences. D3 size and RSF correlate approximately with a morphology-based phylogeny, and within oniscideans RSF increases as more recent divergences occur. The D3 ofArmadillidium vulgare, with an RSF of 1.87, is the highest value recorded for any known expansion segment. Regions of high sequence simplicity in nuclear ribosomal RNA were previously only known from the higher vertebrate lineage. Here we demonstrate that this phenomenon occurs in a more extreme condition within a monophyletic invertebrate lineage. The extreme size changes identified could indicate that expansion segments are an extraneous element in the functioning ribosome.
Similar content being viewed by others
References
Argos P, Rossman MG, Grau UM, Zuber A, Franck G, Tratschin JD (1979) Thermal stability and protein structure. Biochemistry 18:5698–5703
Bernardi G, Bernardi G (1986) Compositional constraints and genome evolution. J Mol Evol 24:1–11
Bernardi G, Olofson B, Filipski B, Zerial M, Salinas J, Cuny G, Meunier-Rotival M, Rodier F (1985) The mosaic genome of warm-blooded vertebrates. Science 228:953–957
Bernardi G, Mouchiroud D, Gautier C, Bernardi G (1988) Compositional patterns in vertebrate genomes: conservation and change in evolution. J Mol Evol 28:7–18
Brusca RC, Wilson GDF (1991) A phylogenetic analysis of the Isopoda with some classificatory recommendations. Mem Qnslnd Museum 31:143–204
Campbell DA, Kubo K, Clark CG, Boothroyd JC (1987) Precise identification of cleavage sites involved in the unusual processing of trypanosome ribosomal RNA. J Mol Biol 196:113–124
Chan Y-L, Olvera J, Wool IG (1983) The structure of rat 28S ribosomal ribonucleic acid inferred from the sequence of nucleotides in a gene. Nucleic Acids Res 11:7819–7831
Clark CG (1987) On the evolution of ribosomal RNA. J Mol Evol 25:343–350
Clark CG, Tague BW, Ware VC, Gerbi SA (1984)Xenopus laevis 28S ribosomal RNA: a secondary structure model and its evolutionary and functional implications. Nucleic Acids Res 12:6197–6220
de Lanversin G, Jacq B (1983) Séquence de la région de la coupure centrale du précurseur de l'ARN ribosomique 26S de Drosophile. C R Acad Sci III 296:1041–1044
de Lanversin G, Jacq B (1989) Sequence and secondary structure of the central domain ofDrosophila 26S rRNA: a universal model for the central domain of the large rRNA containing the region in which the central break may happen. J Mol Evol 28:403–417
Douthwaite S, Christensen A, Garrett RA (1983) Higher order structure in the 3′-minor domain of small subunit ribosomal RNAs from a Gram negative bacterium, a Gram positive bacterium, and a eukaryote. J Mol Biol 169:249–279
Dover GA (1982) Molecular drive: a cohesive mode of species evolution. Nature 299:111–117
Dover GA, Coen E (1981) Springcleaning ribosomal DNA: a model for multigene evolution? Nature 290:731–732
Dover GA, Flavell RB (1984) Molecular co-evolution: rDNA divergence and the maintenance of function. Cell 38:622–623
Edney EB (1960) Terrestrial adaptations. The physiology of Crustacea vol 1. Academic Press, New York, pp 367–393
Edney EB (1968) Transition from water to land in isopod crustaceans. Am Zool 8:309–326
Ellis RE, Sulston JE, Coulson AR (1986) The rDNA ofC. elegans: sequence and structure. Nucleic Acids Res 14:2345–2364
Fujiwara H, Ishikawa H (1986) Molecular mechanism of introduction of the hidden break into the 28S rRNA of insects: implication based on structural studies. Nucleic Acids Res 14:6393–6401
Gerbi SA (1985) Evolution of ribosomal RNA. In: MacIntyre RJ (ed) Molecular evolutionary genetics. Plenum, New York, pp 419–517
Gonzalez IL, Gorski JL, Campen TJ, Dorney DJ, Erickson JM, Sylvester JE, Schmickel RD (1985) Variation among human 28S ribosomal RNA genes. Proc Natl Acad Sci USA 82:7666–7670
Gutell RR, Schnare MN, Gray, MW (1990) A compilation of large subunit (23S-like) ribosomal RNA sequences presented in a secondary structure format. Nucleic Acids Res [Suppl] 18:r2319-r2330
Gyllensten UB, Erlich HA (1988) Generation of single-stranded DNA by the polymerase chain reaction and its application to direct sequencing of theHLA-DQA locus. Proc Natl Acad Sci USA 85:7652–7656
Hancock JM, Armstrong, JS (1994) SIMPLE34: an improved and enhanced implementation for VAX and Sun computers of the SIMPLE algorithm for analysis of clustered repetitive motifs in nucleotide sequences. Comp Appl Biosci 10:67–70
Hancock JM, Dover GA (1988) Molecular coevolution among cryptically simple expansion segments of eukaryotic 26S/28S rRNAs. Mol Biol Evol 5:377–391
Hancock JM, Dover GA (1990) ‘Compensatory slippage’ in the evolution of ribosomal RNA genes. Nucleic Acids Res 18:5949–5954
Hassouna N, Michot B, Bachellerie J-P (1984) The complete nucleotide sequence of mouse 28S rRNA gene. Implications for the process of size increase of the large subunit rRNA in higher eukaryotes. Nucleic Acids Res 12:3563–3583
Larson A (1991) Evolutionary analysis of length-variable sequences: divergent domains of ribosomal RNA. In: Miyamoto MM, Cracraft J (eds) Phylogenetic analysis of DNA sequences. Oxford University Press, pp 221–248
Larson A, Wilson AC (1989) Patterns of ribosomal RNA evolution in salamanders. Mol Biol Evol 6:131–154
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning. A laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Michot B, Bachellerie J-P (1987) Comparisons of large sub-unit rRNAs reveal some eukaryote-specific elements of secondary structure. Biochimie 69:11–23
Michot B, Qu L-H, Bachellerie J-P (1990) Evolution of large sub-unit rRNA structure. The diversification of divergent D3 domain among major phylogenetic groups. Eur J Biochem 188:219–229
Nelles L, Van Broeckhoven C, de Wachter R, Vandenberghe A (1984) Location of the hidden break in large subunit ribosomal RNA ofArtemia salina. Naturwissenschaften 71:634–635
Noller HF (1984) Structure of ribosomal RNA. Annu Rev Biochem 53:119–162
Peattie DA, Douthwaite S, Garrett RA, Noller HF (1981) A ‘bulged’ double helix in a RNA-protein contact site. Proc Natl Acad Sci USA 78:7331–7335
Rousset F, Pélandakis M, Solignac M (1991) Evolution of compensatory substitutions through G · U intermediate state inDrosophila rRNA. Proc Natl Acad Sci USA 88:10032–10036
Ruiz Linares AR, Hancock JM, Dover GA (1991) Secondary structure constraints on the evolution ofDrosophila 28S ribosomal RNA expansion segments. J Mol Biol 219:381–390
Salinas J, Matassi G, Montero LM, Bernardi G (1988) Compositional compartmentalisation and compositional patterns in the nuclear genomes of plants. Nucleic Acids Res 16:4269–4285
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467
Schmalfuss H (1989) Phylogenetics in Oniscidea. Monitore Zool Ital Monograph 4:3–27
Sueoka N (1988) Directional mutation pressure and neutral molecular evolution. Proc Natl Acad Sci USA 85:2653–2657
Tautz D, Trick M, Dover GA (1986) Cryptic simplicity in DNA is a major source of genetic variation. Nature 322:652–656
Tautz D, Hancock JM, Webb DA, Tautz C, Dover GA (1988) Complete sequences of the rRNA genes ofDrosophila melanogaster. Mol Biol Evol 5:366–376
Wada A, Suyama A (1986) Local stability of DNA and RNA secondary structure and its relation to biological function. Prog Biophys Mol Biol 47:113–157
Ware VC, Tague BW, Clark CG, Gourse RL, Brand RC, Gerbi SA (1983) Sequence analysis of 28S ribosomal DNA from the amphibianXenopus laevis. Nucleic Acids Res 11:7795–7817
Ware VC, Renkawitz R, Gerbi SA (1985) rRNA processing: removal of only nineteen bases at the gap between 28S∂ and 28Sβ rRNAs inSciara coprophila. Nucleic Acids Res 13:3581–3597
Woese CR, Gutell R, Gupta R, Noller HF (1983) A detailed analysis of the higher order structure of 16S-like ribosomal ribonucleic acids. Microbiol Rev 47:621–669
Zuker M, Stiegler P (1981) Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res 9:133–148
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Nunn, G.B., Theisen, B.F., Christensen, B. et al. Simplicity-correlated size growth of the nuclear 28S ribosomal RNA D3 expansion segment in the crustacean order isopoda. J Mol Evol 42, 211–223 (1996). https://doi.org/10.1007/BF02198847
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02198847